|
The ligation procedure was carried out at 37☌ for 3 h to generate DNA template adapter. Genomic DNA (100 ng/μL) was digested with Eco RI (5 U/μL) (Boehringer Mannheim GmbH, Germany) and Tru 9I (5 U/μL) (Roche Diagnostics GmbH, Germany) in buffer A (×10) (Promega, USA) at 37☌ for 1 h. The results might provide some useful information for studying the genetic relationship of the genus Curcuma.Īmplified fragment length polymorphism fingerprintingĪFLP fingerprinting was performed as described by Vos et al., 1995, with some modification. As a result, this study is aimed to evaluate the phylogenetic relationships of twenty Curcuma species existing in Thailand, using the AFLP marker. Among this, AFLP not only has higher reproducibility and no prior sequence information but also has the capability to detect various polymorphisms in the genome simultaneously. Polymerase chain reaction (PCR) based on molecular markers was used to support the identification and distinguishing the genetic diversity analysis in medicinal plants such as the simple sequence repeat, random amplified polymorphic DNA (RAPD), and amplified fragment length polymorphism (AFLP) technique. Recently, several molecular techniques have been employed for taxonomic identification. 
Moreover, in some cases, during the early flowering stage, the similarities in morphology led to confusion in their identification. 
The identification of Curcuma species has not yet been accomplished as there are some main problems in the taxonomic studies and lack of type specimens, etc. The diarylheptanoids are the main active components in the rhizomes of Curcuma plants. In addition, the Curcuma species was used in folk medicine for the treatment of diarrhea, dysentery, bronchial complaints, pneumonia, insect bites, and infectious wounds. 
Several Curcuma species have long been known for their use as food, spices, cosmetics, and ornamental plants. Plants in the genus Curcuma (family Zingiberaceae) comprise of >100 species widely distributed in Asia-tropical and Asia-Pacific regions. In summary, the ten successful AFLP primer combinations could be used to determine the genetic relationship among closely related twenty Curcuma species in Thailand. Cluster III belongs to Curcuma singularis and Alpinia galanga (outgroup plant), which clearly separated into different clusters from twenty Curcuma species. Cluster II can be subdivided into IIA being composed of Curcuma longa, Curcuma Zedoaria, and Curcuma aromatica with the SI 0.8989–0.9071, whereas Cluster IIB was composed of Curcuma leucorrhiza, Curcuma aeruginosa, Curcuma comosa, Curcuma mangga, Curcuma angustifolia, Curcuma amada, Curcuma sessilis, and Curcuma albicoma with the SI 0.8236–0.9500. Cluster I can be subdivided into IA, which composed of Curcuma parviflora, Curcuma sparganiifolia, Curcuma alismatifolia, Curcuma larsenii, Curcuma Gracillima, and Curcuma rhabdota with similarity index (SI) 0.7926–0.9358 and IB composed of Curcuma petiolata and Curcuma rubrobracteata with the SI 0.9240. The dendrogram generated from the unweighted pair group method of the arithmetic average could separate these Curcuma species into three major clusters. AFLP fingerprint showed 98.54% highly polymorphisms with the number of bands (617 bands) ranging between 48 and 80 bands. In this study, the genetic relationship among twenty Curcuma species from Thailand was accessed by the amplified fragment length polymorphism (AFLP) method. 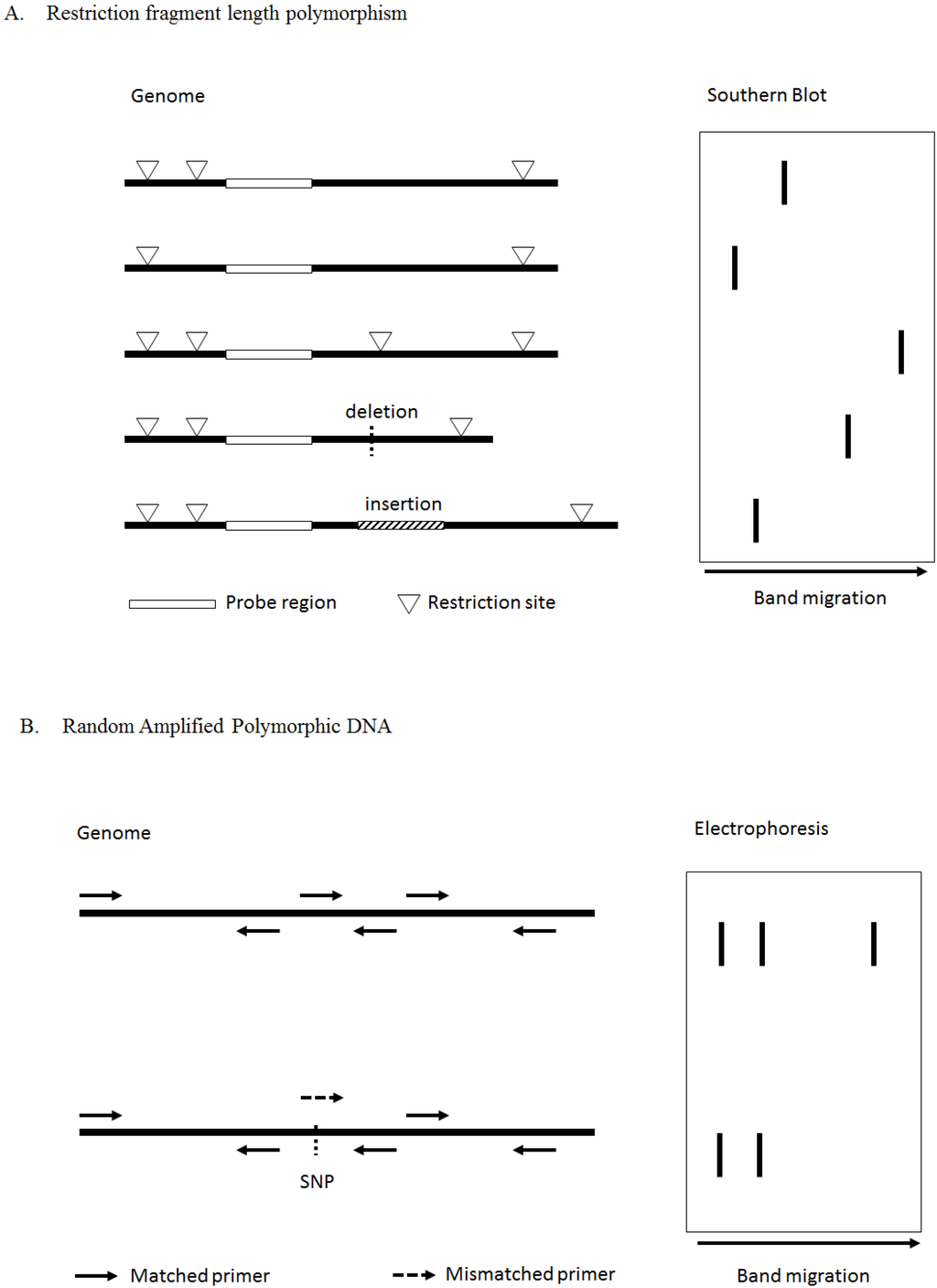
Recently, the molecular technique is one of the reliable and powerful tools for plant identification. However, the identification of plants in the genus Curcuma is very difficult due to morphological similarity in the early flowering stage. It has long been known for their uses as folk medicines, foods, spices, and cosmetics. Plants in the genus Curcuma are a rhizomatous perennial herb which is widely distributed in Thailand.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
